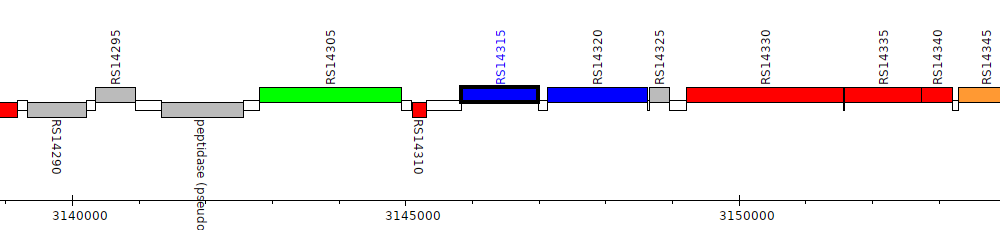
Burkholderia cenocepacia AU 1054, BCEN_RS14315
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071973 | bacterial-type flagellum-dependent cell motility |
Inferred from Sequence Model
Term mapped from: InterPro:PF00700
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005198 | structural molecule activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00700
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009420 | bacterial-type flagellum filament |
Inferred from Sequence Model
Term mapped from: InterPro:PR00207
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF64518 | 1 | 384 | 1.44E-90 | |||
| Gene3D | G3DSA:1.20.1330.10 | 43 | 333 | 5.3E-66 | |||
| Coils | Coil | 301 | 321 | - | |||
| PRINTS | PR00207 | Flagellin signature | IPR001492 | Flagellin | 36 | 53 | 6.6E-24 |
| PRINTS | PR00207 | Flagellin signature | IPR001492 | Flagellin | 110 | 133 | 6.6E-24 |
| Pfam | PF00700 | Bacterial flagellin C-terminal helical region | IPR001029 | Flagellin, D0/D1 domain | 300 | 383 | 2.2E-18 |
| Pfam | PF00669 | Bacterial flagellin N-terminal helical region | IPR001029 | Flagellin, D0/D1 domain | 4 | 140 | 3.1E-39 |
| Gene3D | G3DSA:1.10.8.410 | IPR042187 | Flagellin, C-terminal domain, subdomain 2 | 334 | 384 | 2.1E-20 | |
| Pfam | PF12613 | Flagellin structural protein | IPR022578 | Flagellin, domain of unknown function | 238 | 290 | 2.5E-24 |
| Gene3D | G3DSA:3.30.70.2120 | 172 | 282 | 5.3E-66 | |||
| PRINTS | PR00207 | Flagellin signature | IPR001492 | Flagellin | 75 | 94 | 6.6E-24 |
| Coils | Coil | 380 | 384 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.