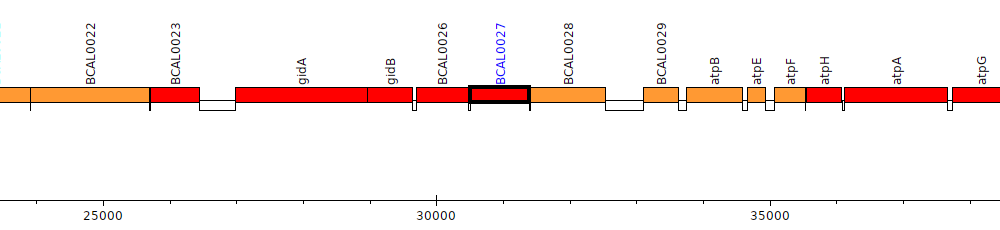
Burkholderia cenocepacia J2315, BCAL0027
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00180
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00470 | IPR003115 | ParB/Sulfiredoxin | 35 | 125 | 2.8E-34 | |
| Pfam | PF17762 | HTH domain found in ParB protein | IPR041468 | ParB/Spo0J, HTH domain | 176 | 224 | 2.4E-17 |
| Coils | Coil | 279 | 297 | - | |||
| CDD | cd16393 | SPO0J_N | 36 | 130 | 6.19326E-51 | ||
| TIGRFAM | TIGR00180 | parB_part: ParB/RepB/Spo0J family partition protein | IPR004437 | ParB/RepB/Spo0J partition protein | 35 | 207 | 3.1E-58 |
| SUPERFAMILY | SSF109709 | 118 | 224 | 3.66E-35 | |||
| Coils | Coil | 108 | 128 | - | |||
| Gene3D | G3DSA:1.10.10.2830 | 112 | 225 | 4.9E-38 | |||
| Pfam | PF02195 | ParB-like nuclease domain | IPR003115 | ParB/Sulfiredoxin | 39 | 125 | 2.9E-26 |
| Gene3D | G3DSA:3.90.1530.30 | 48 | 111 | 1.6E-28 | |||
| SUPERFAMILY | SSF110849 | IPR036086 | ParB/Sulfiredoxin superfamily | 37 | 127 | 1.33E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.